Tree of the month: Neurospora crassa SO gene and hyphal fusion
 Since March the town of Saint-Pièrre, in Martinique Island on the Caribbean sea, was observing the timid increase in activity of Mount Pelèe, a volcano six and a half kilometers apart. It all started with some fumaroles, that revealed the mountain was awaking. At the beginning of May the volcano started to spit clouds of ashes. Finally, during the morning of 8th of May of 1902, all communications with town, suddenly ceased. The volcano erupted, and the town was engulfed by a black cloud of scorching gas and ashes. Nearly all the 30,000 inhabitants of Saint Pierre perished instantaneously, and chroniques of the tragedy report that one of the only survivors was a pity criminal that had the fortune of being arrested in an underground cell.
Since March the town of Saint-Pièrre, in Martinique Island on the Caribbean sea, was observing the timid increase in activity of Mount Pelèe, a volcano six and a half kilometers apart. It all started with some fumaroles, that revealed the mountain was awaking. At the beginning of May the volcano started to spit clouds of ashes. Finally, during the morning of 8th of May of 1902, all communications with town, suddenly ceased. The volcano erupted, and the town was engulfed by a black cloud of scorching gas and ashes. Nearly all the 30,000 inhabitants of Saint Pierre perished instantaneously, and chroniques of the tragedy report that one of the only survivors was a pity criminal that had the fortune of being arrested in an underground cell.
In nature, the scope of catastrophic events goes far beyond hurricanes and volcanoes; and depending on your scale it may be something as mundane as a drying pond or some days of unusual temperatures. Catastrophes cause an indiscriminate loss of population size, where luck instead of some phenotypic quality is what defines the survivors. Sudden population loss causes the phenomenon known as “population bottleneck” that produces a rather low genetic diversity in the population. This phenomenon is due to an initial existence of a low number of individuals and has important biological consequences. For instance, population bottlenecks is a common factor limiting the recovering of endangered species. However, while catastrophes itself are mostly a deadly lottery, some organisms have adapted to win it every time. Organisms with a fast life cycle and the ability to quickly exploit a wide range of food sources greatly benefit the sudden loss of best fitted competitors, and are usually the first colonizers after a cataclysm.
 For example, not long after a fire in a forest is quite common to see an orange and fluffy organisms sprouting from the piles of burned wood. This pioneer is a humble mould known as Neurospora crassa. It is a generalistic saprotrophic, and one of the filamentous fungi with fastest growth ratio. As its spores are tiny and can travel big distances transported by the wind, Neurospora has no problem arriving to an area that has been barren of most lifeforms. Then, using its fast growth it can outcompete other molds, at least while the food is abundant and easy to obtain. Thanks to this fast growth, Neurospora crassa was able to contaminate and propagate in french bakeries during the XIX century, from where it was first isolated. Years later, its fast growth and well known spore development made Neurospora the organism of choice of Edward Tatum and George Wells Beadle, who used our microscopic friend to prove that DNA control essential biochemical pathways, a work justly awarded with a Nobel prize in 1958.
For example, not long after a fire in a forest is quite common to see an orange and fluffy organisms sprouting from the piles of burned wood. This pioneer is a humble mould known as Neurospora crassa. It is a generalistic saprotrophic, and one of the filamentous fungi with fastest growth ratio. As its spores are tiny and can travel big distances transported by the wind, Neurospora has no problem arriving to an area that has been barren of most lifeforms. Then, using its fast growth it can outcompete other molds, at least while the food is abundant and easy to obtain. Thanks to this fast growth, Neurospora crassa was able to contaminate and propagate in french bakeries during the XIX century, from where it was first isolated. Years later, its fast growth and well known spore development made Neurospora the organism of choice of Edward Tatum and George Wells Beadle, who used our microscopic friend to prove that DNA control essential biochemical pathways, a work justly awarded with a Nobel prize in 1958.
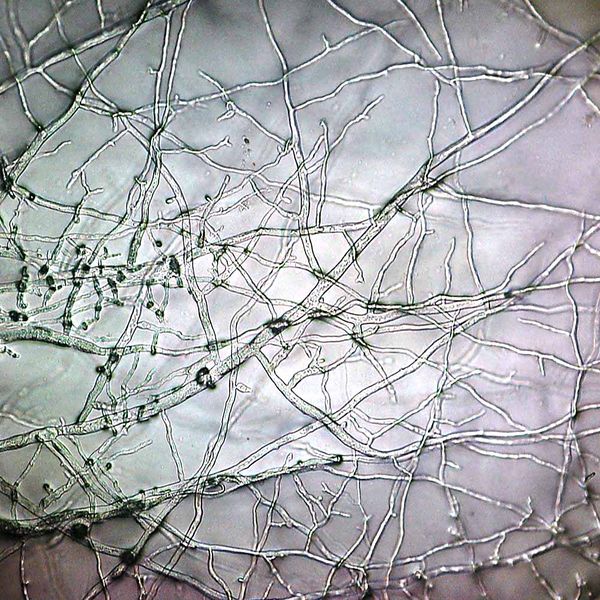 Fungal hyphae can extend over surprisingly large distances, distributing the nutrients they obtain across the fungus by complex hydraulic systems. Such hydraulic engines are also behind Neurospora’s trick for achieving such a fast growth. There is a limit to the speed at which eukaryotic nuclei can divide, so in order to fill the nascent hyphae at such speed Neurospora would need to multiply the nuclei at its tip at a rate that would defy any logic. Instead, most of the mycelia is producing new nuclei that are transported to the expanding tip by cytoplasmic flux. Naturally, since nutrient flux usually goes from the tip to the rest of the hyphae, nuclear flux goes in the opposite direction, and both movement are mediated by raw liquid flow; the hyphae must be organized in an interconnected network, with some parts of the fungus acting as roadways in one sense and other in the opposite. Such organization requires secondary transversal hyphae interconnecting the system. It is very similar to the in the way human roads expand and interconnect across the landscape in response to demographic and economical factors. This interconnections are formed by the fusion between two hyphae. Neurospora mutants of the gene SOFT are unable to perform this hyphal fusion, growing in an ever branching way, akin to the roots of a tree.
Fungal hyphae can extend over surprisingly large distances, distributing the nutrients they obtain across the fungus by complex hydraulic systems. Such hydraulic engines are also behind Neurospora’s trick for achieving such a fast growth. There is a limit to the speed at which eukaryotic nuclei can divide, so in order to fill the nascent hyphae at such speed Neurospora would need to multiply the nuclei at its tip at a rate that would defy any logic. Instead, most of the mycelia is producing new nuclei that are transported to the expanding tip by cytoplasmic flux. Naturally, since nutrient flux usually goes from the tip to the rest of the hyphae, nuclear flux goes in the opposite direction, and both movement are mediated by raw liquid flow; the hyphae must be organized in an interconnected network, with some parts of the fungus acting as roadways in one sense and other in the opposite. Such organization requires secondary transversal hyphae interconnecting the system. It is very similar to the in the way human roads expand and interconnect across the landscape in response to demographic and economical factors. This interconnections are formed by the fusion between two hyphae. Neurospora mutants of the gene SOFT are unable to perform this hyphal fusion, growing in an ever branching way, akin to the roots of a tree.
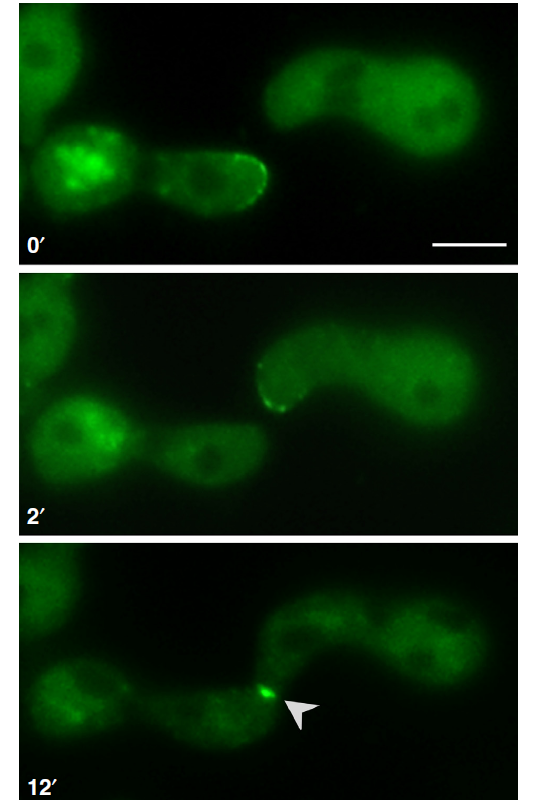 As we can see in our tree of the month, that is part of the metaphylome of fungal model species, SOFT appears in several unrelated pezizomycotina with a very stable evolutionary history. Looking at an additional tree with more species, such as this example from the Fusarium oxysporum phylome, reveals that the pattern is much more extended. The gene SOFT is conserved across all filamentous ascomycota groups (Pezizomycotina), with very few duplication events. It is absent in ascomycetous yeasts (for instance in Saccharomycotina and in Schizosaccharomyces), and in basidiomycota, suggesting that the network organization has significant differences between these two major groups of fungi. Little is known about the physiology of this protein, but it is predicted to interact with other proteins, virtue of the presence of some protein-protein interaction domains, and can be localized in vivo in septal plugs, structures that act as gates between compartments in the hyphae. In the picture next to this paragraph you can observe cell fusion of Neurospora crasa cells in which SO protein has been labelled using GFP. During hyphal fusion, the two fusing hyphae seem to enter in a molecular conversation, alternating between a receptor and an emissor state, one of which is characterized by the expression of SO. Observe that the expression of SO is high in the right cell of the picture in the first pannel and low in the second; while for the left cell the pattern is the opposite. SO finally accumulates in the new anastomosis, as we can observe in the third pannel of the picture. Cell fusion is also important for sexual reproduction in filamentous fungi, and consequently Neurospora mutants for SO cannot mate.
As we can see in our tree of the month, that is part of the metaphylome of fungal model species, SOFT appears in several unrelated pezizomycotina with a very stable evolutionary history. Looking at an additional tree with more species, such as this example from the Fusarium oxysporum phylome, reveals that the pattern is much more extended. The gene SOFT is conserved across all filamentous ascomycota groups (Pezizomycotina), with very few duplication events. It is absent in ascomycetous yeasts (for instance in Saccharomycotina and in Schizosaccharomyces), and in basidiomycota, suggesting that the network organization has significant differences between these two major groups of fungi. Little is known about the physiology of this protein, but it is predicted to interact with other proteins, virtue of the presence of some protein-protein interaction domains, and can be localized in vivo in septal plugs, structures that act as gates between compartments in the hyphae. In the picture next to this paragraph you can observe cell fusion of Neurospora crasa cells in which SO protein has been labelled using GFP. During hyphal fusion, the two fusing hyphae seem to enter in a molecular conversation, alternating between a receptor and an emissor state, one of which is characterized by the expression of SO. Observe that the expression of SO is high in the right cell of the picture in the first pannel and low in the second; while for the left cell the pattern is the opposite. SO finally accumulates in the new anastomosis, as we can observe in the third pannel of the picture. Cell fusion is also important for sexual reproduction in filamentous fungi, and consequently Neurospora mutants for SO cannot mate.
References:
http://www.ncbi.nlm.nih.gov/pubmed/26226449
http://www.ncbi.nlm.nih.gov/pubmed/16079346
http://www.ncbi.nlm.nih.gov/pubmed/21640311
http://www.ncbi.nlm.nih.gov/pubmed/26340439
http://www.ncbi.nlm.nih.gov/pubmed/11988765
http://www.ncbi.nlm.nih.gov/pubmed/24875794
Pictures:
Neurospora on burned tree picture from Namboori B. Raju, Stanford University
Mycelium picture from Bob Blaylock
GFP labelling of SO protein during hyphal fusion in Neurospora extracted from Herzog, Schumann & Fleißner (2015)
